R1b Potts (R-M269)
R1b – Eastern European Potts Family Line
Haplogroup R1b1a2 (R-M269)
Over 30 samples of POTTS Ancestry have had yDNA testing and have been confirmed as haplogroup R-M269 making it by far the most prolific bunch represented in the available samples. It would be good to get further subclade testing done on these pots so as to see how they look further down the phylogenetic tree. Are the R-L21 (R1b1a2a1a2c)? At least one member is. It would be good to know if the majority are as well.
R1b Potts Cluster 1
- Samuel Potts, 1789, Iredell Co., North Carolina
- Henry Potts 1 of Iredell Co.,North Carolina
- Stephen Potts fr John Potts 1 of Iredell North Carolina
- James McCuen Potts North Carolina – Born 1815
- Robert Potts, b 1771 in North Carolina (from John 1??)
- John 1 Potts of Iredell County, North Carolina
- William Howard Potts – 1828 of Georgia
R1b Potts Cluster 2
- Thomass Pott Llangirrig, Montgomerys
- Lewis Smith Martin
- John Potts – 1760-1839 – Oldham County, Kentucky
- John Potts
more to come . . .
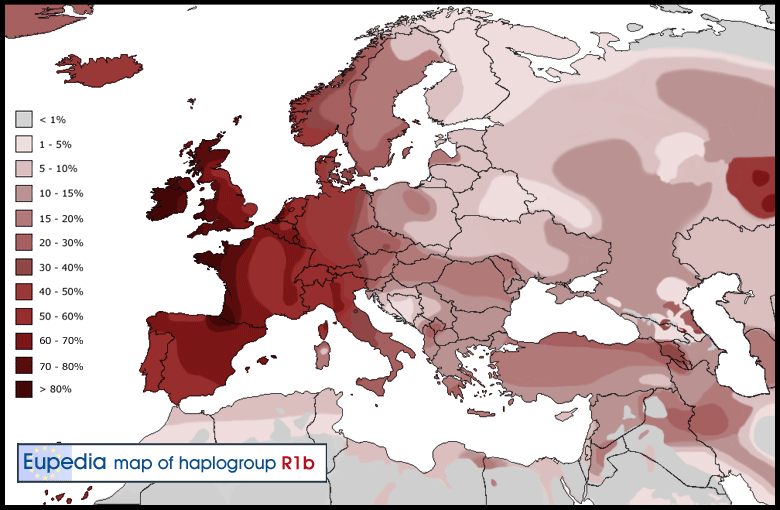
R1b is the most common haplogroup in Western Europe, reaching over 80% of the population in Ireland, the Scottish Highlands, western Wales, the Atlantic fringe of France, the Basque country and Catalonia. It is also common in Anatolia and around the Caucasus, in parts of Russia and in Central and South Asia. Besides the Atlantic and North Sea coast of Europe, hotspots include the Po valley in north-central Italy (over 70%), Armenia (35%), the Bashkirs of the Urals region of Russia (50%), Turkmenistan (over 35%), the Hazara people of Afghanistan (35%), the Uyghurs of North-West China (20%) and the Newars of Nepal (11%). R1b-V88, a subclade specific to sub-Saharan Africa, is found in 60 to 95% of men in northern Cameroon.
The frequency is about 92% in Wales, 82% in Ireland, 70% in Scotland, 68% in Spain, 60% in France (76% in Normandy), about 60% in Portugal, 53% in Italy,[7] 45% in Eastern England, 50% in Germany, 50% in the Netherlands, 42% in Iceland, and 43% in Denmark. It is as high as 95% in parts of Ireland.
In articles published around 2000 it was proposed that this clade had been in Europe before the last ice age,[43] but by 2010 more recent periods such as the European Neolithic have become the focus of proposals. A range of newer estimates for R1b1a2, or at least its dominant parts in Europe, are from 4,000 to a maximum of about 10,000 years ago, and looking in more detail is seen as suggesting a migration from Western Asia via southeastern Europe.[1][7][12][17]
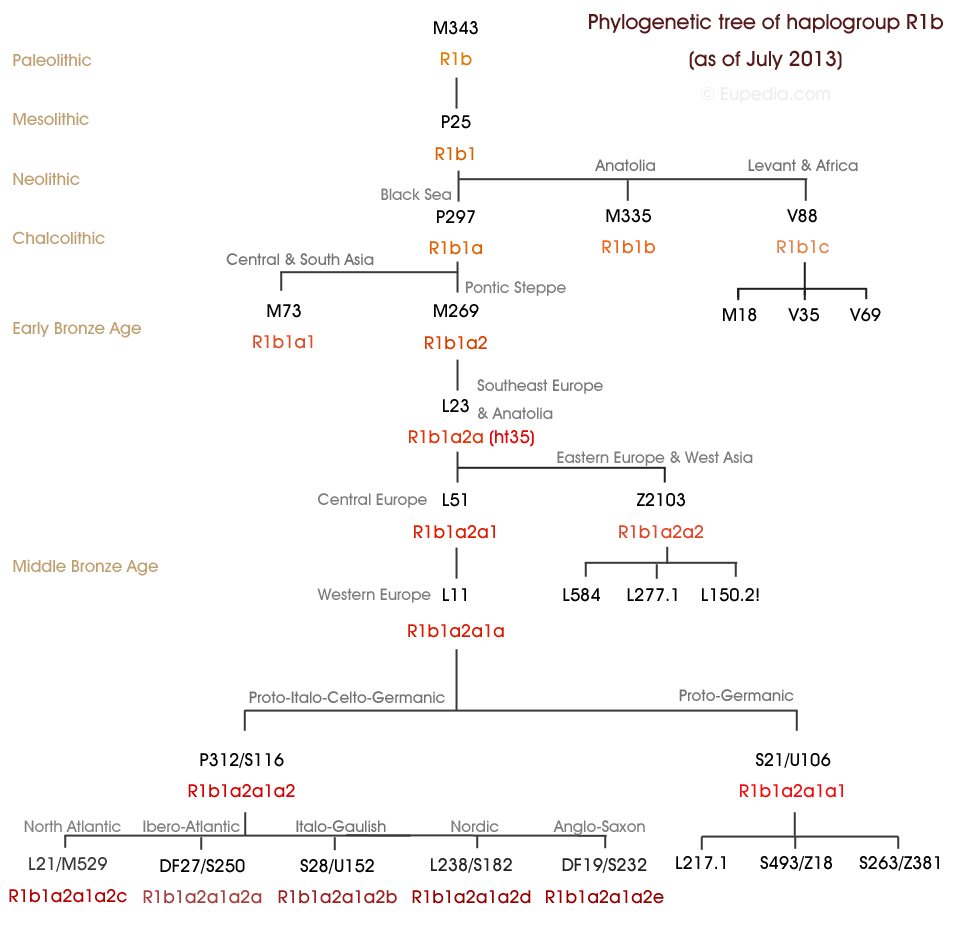
R-M269 or L21 History
Add more history here later once confirmed if L21 or not.
Resources
- Eupedia – Haplogroup R1b
- Wikipedia – Haplogroup R1b
Glossary
- Clade comes from the Greek word Klados = branch. A Clade on the Y Chromosome tree is also called a Haplogroup. Subclade is a term to describe the relationship between two clades with the sub-clade being downstream (occurring later in time). A Clade includes all the descendants of a single founder (common ancestor).
- Haplogroup is defined as a group of similar Haplotypes that share a common ancestor with a Single Nucleotide Polymorphism (SNP) mutation. Your Haplogroup tells much about your ancient ethnic origins. Meaning thousands of years ago. In the case of our I2a Haplogroup, we all share the SNP (called Snip) I-P37.2+. The I2a Haplogroup has numerous subgroups or subclades, all determined by various other SNPs which we will discuss later.
- SNP is defined as a change in the DNA which happens when a single nucleotide (A, T, G or C) in the genome sequence is altered. A person has many SNPs that together create a unique DNA pattern for that individual. Snips clarify the branching of a tree–separation of different subhaplogroups.
- Haplotype is defined as one person’s set of values for the DYS markers that have been tested. In other words, your yDNA test results. Think of Haplotypes as leaves on a tree, and a Haplogroup as a limb of that tree. Haplotype is a contaction of the phrase “haploid genotype”.
- Allele is defined as a DNA sequence that repeats at a certain location (DYS marker) on the Y Chromosome. The Allele value is the number of times the sequence repeats. Pronounced uh-LEEL.
- STR is a short DNA motif (pattern) repeated in tandem. A, T, G or C repeated eleven times would give the DYS marker a value or allele of eleven. DYS is short for DNA/ Y-Chromosome/ Segment. The name of a marker on the Y-Chromosome.